Nearest Neighbor Projection is an iterative method for visualizing high-dimensional dataset
in that a data is sequentially located in the low-dimensional space by maintaining
the triangular distance spread of target data with its two nearest neighbors in the high-dimensional space.
We extended the original method to be applied for arbitrarily low-dimensional space. Due the generalization,
we opted for a global optimization method of Differential Evolution (DEoptim) within in that it may add computational burden to certain degrees.
do.nnp(
X,
ndim = 2,
preprocess = c("null", "center", "scale", "cscale", "whiten", "decorrelate")
)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
References
Tejada E, Minghim R, Nonato LG (2003). “On Improved Projection Techniques to Support Visual Exploration of Multidimensional Data Sets.” Information Visualization, 2(4), 218–231.
Examples
# \donttest{
## use iris data
data(iris)
set.seed(100)
subid = sample(1:150,50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
## let's compare with other methods
out1 <- do.nnp(X, ndim=2) # NNP
out2 <- do.pca(X, ndim=2) # PCA
out3 <- do.dm(X, ndim=2) # Diffusion Maps
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
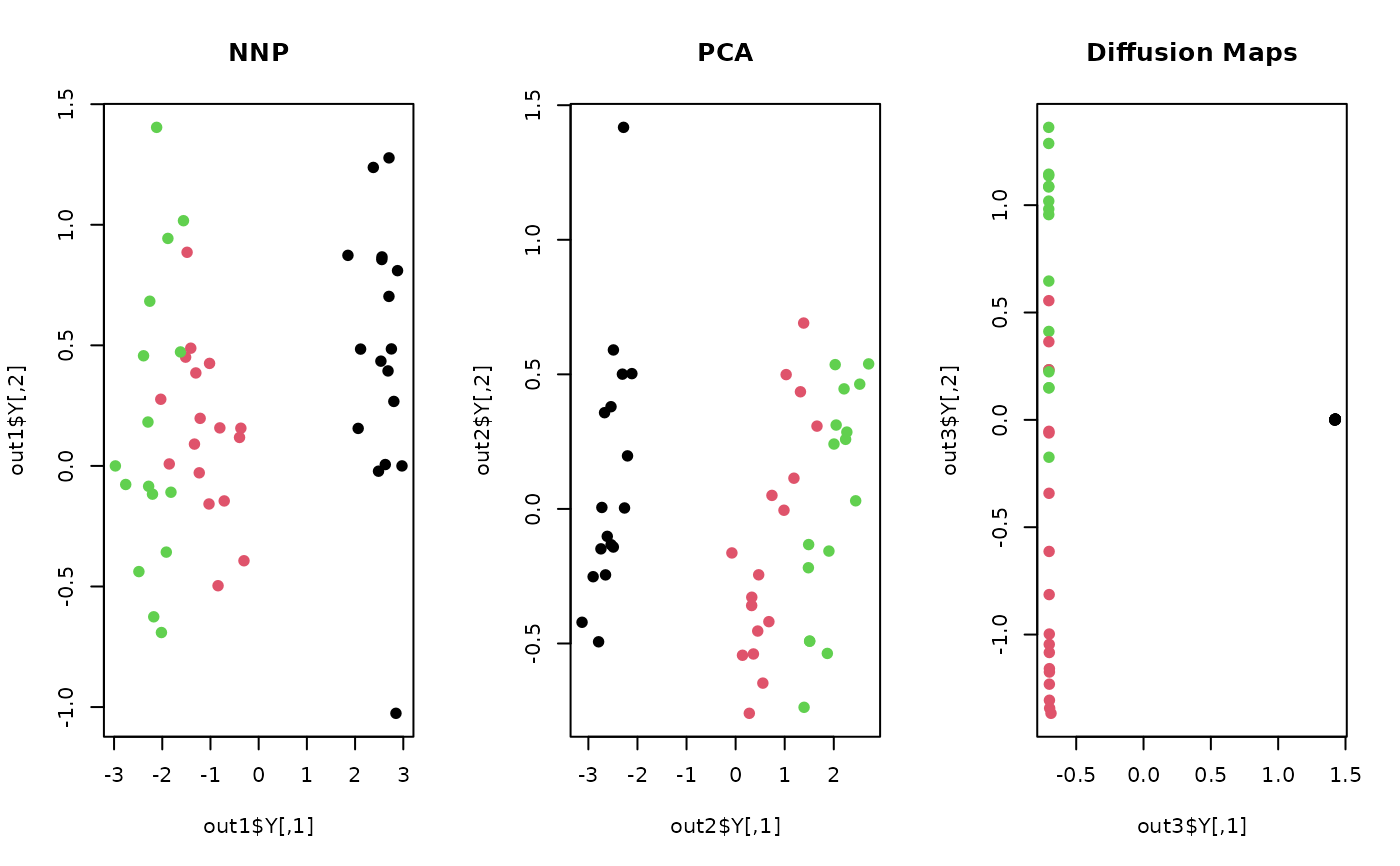
plot(out1$Y, pch=19, col=label, main="NNP")
plot(out2$Y, pch=19, col=label, main="PCA")
plot(out3$Y, pch=19, col=label, main="Diffusion Maps")
 par(opar)
# }
par(opar)
# }