do.sammon is an implementation for Sammon mapping, one of the earliest
dimension reduction techniques that aims to find low-dimensional embedding
that preserves pairwise distance structure in high-dimensional data space.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.- initialize
"random"or"pca"; the former performs fast random projection (see alsodo.rndproj) and the latter performs standard PCA (see alsodo.pca).
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
References
Sammon, J.W. (1969) A Nonlinear Mapping for Data Structure Analysis. IEEE Transactions on Computers, C-18 5:401-409.
Sammon JW (1969). “A Nonlinear Mapping for Data Structure Analysis.” IEEE Transactions on Computers, C-18(5), 401–409.
Examples
# \donttest{
## load iris data
data(iris)
X = as.matrix(iris[,1:4])
label = as.factor(iris$Species)
## compare two initialization
out1 = do.sammon(X,ndim=2) # random projection
out2 = do.sammon(X,ndim=2,initialize="pca") # pca as initialization
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,2))
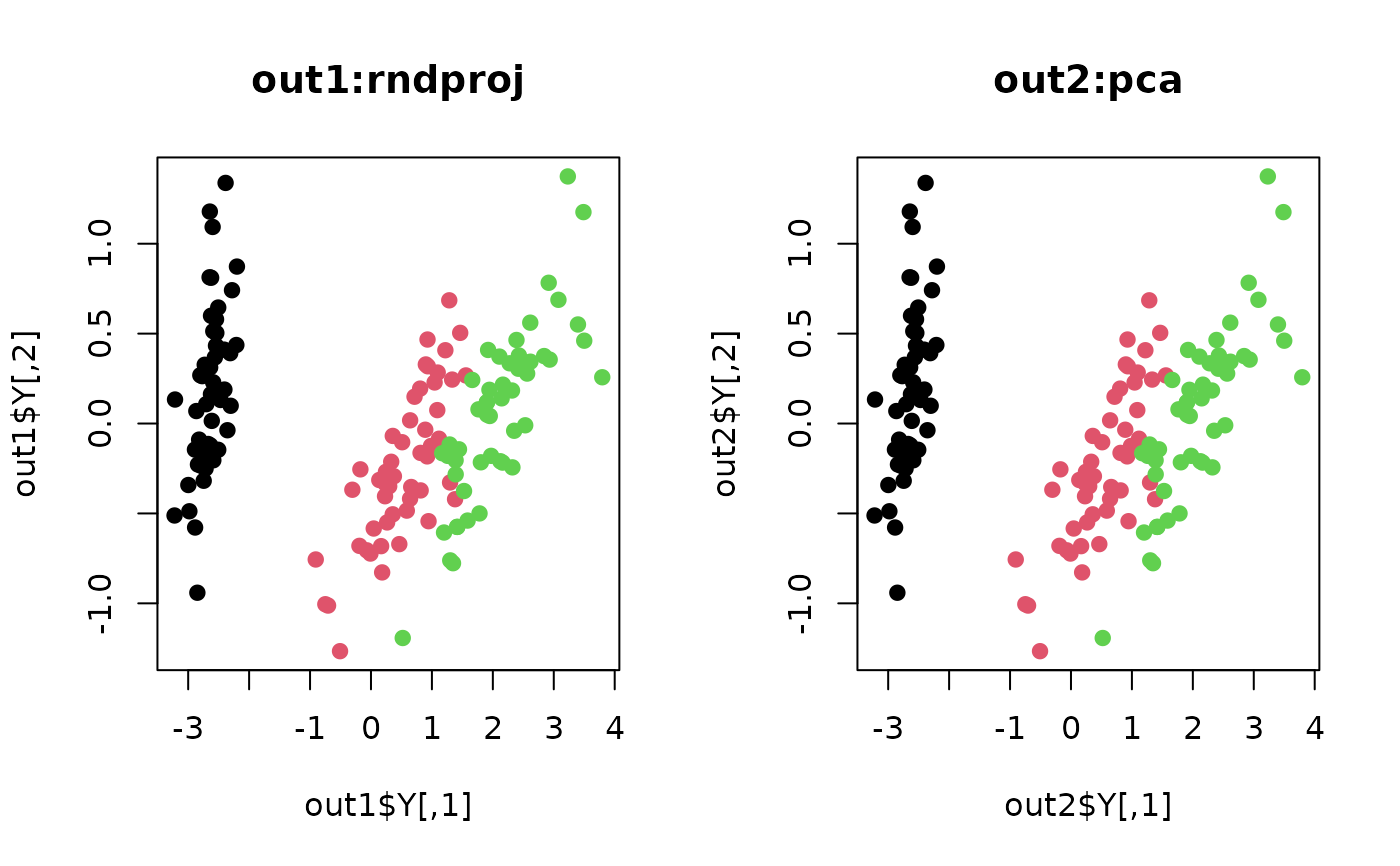
plot(out1$Y, pch=19, col=label, main="out1:rndproj")
plot(out2$Y, pch=19, col=label, main="out2:pca")
 par(opar)
# }
par(opar)
# }