Sparse PCA (do.spca) is a variant of PCA in that each loading - or, principal
component - should be sparse. Instead of using generic optimization package,
we opt for formulating a problem as semidefinite relaxation and utilizing ADMM.
do.spca(X, ndim = 2, mu = 1, rho = 1, ...)Arguments
- X
an \((n\times p)\) matrix whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- mu
an augmented Lagrangian parameter.
- rho
a regularization parameter for sparsity.
- ...
extra parameters including
- maxiter
maximum number of iterations (default: 100).
- abstol
absolute tolerance stopping criterion (default: 1e-8).
- reltol
relative tolerance stopping criterion (default: 1e-4).
Value
a named Rdimtools S3 object containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
- algorithm
name of the algorithm.
References
Zou H, Hastie T, Tibshirani R (2006). “Sparse Principal Component Analysis.” Journal of Computational and Graphical Statistics, 15(2), 265–286.
d'Aspremont A, El Ghaoui L, Jordan MI, Lanckriet GRG (2007). “A Direct Formulation for Sparse PCA Using Semidefinite Programming.” SIAM Review, 49(3), 434–448.
Ma S (2013). “Alternating Direction Method of Multipliers for Sparse Principal Component Analysis.” Journal of the Operations Research Society of China, 1(2), 253–274.
See also
Examples
# \donttest{
## use iris data
data(iris, package="Rdimtools")
set.seed(100)
subid = sample(1:150,50)
X = as.matrix(iris[subid,1:4])
lab = as.factor(iris[subid,5])
## try different regularization parameters for sparsity
out1 <- do.spca(X,ndim=2,rho=0.01)
out2 <- do.spca(X,ndim=2,rho=1)
out3 <- do.spca(X,ndim=2,rho=100)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
plot(out1$Y, col=lab, pch=19, main="SPCA::rho=0.01")
plot(out2$Y, col=lab, pch=19, main="SPCA::rho=1")
plot(out3$Y, col=lab, pch=19, main="SPCA::rho=100")
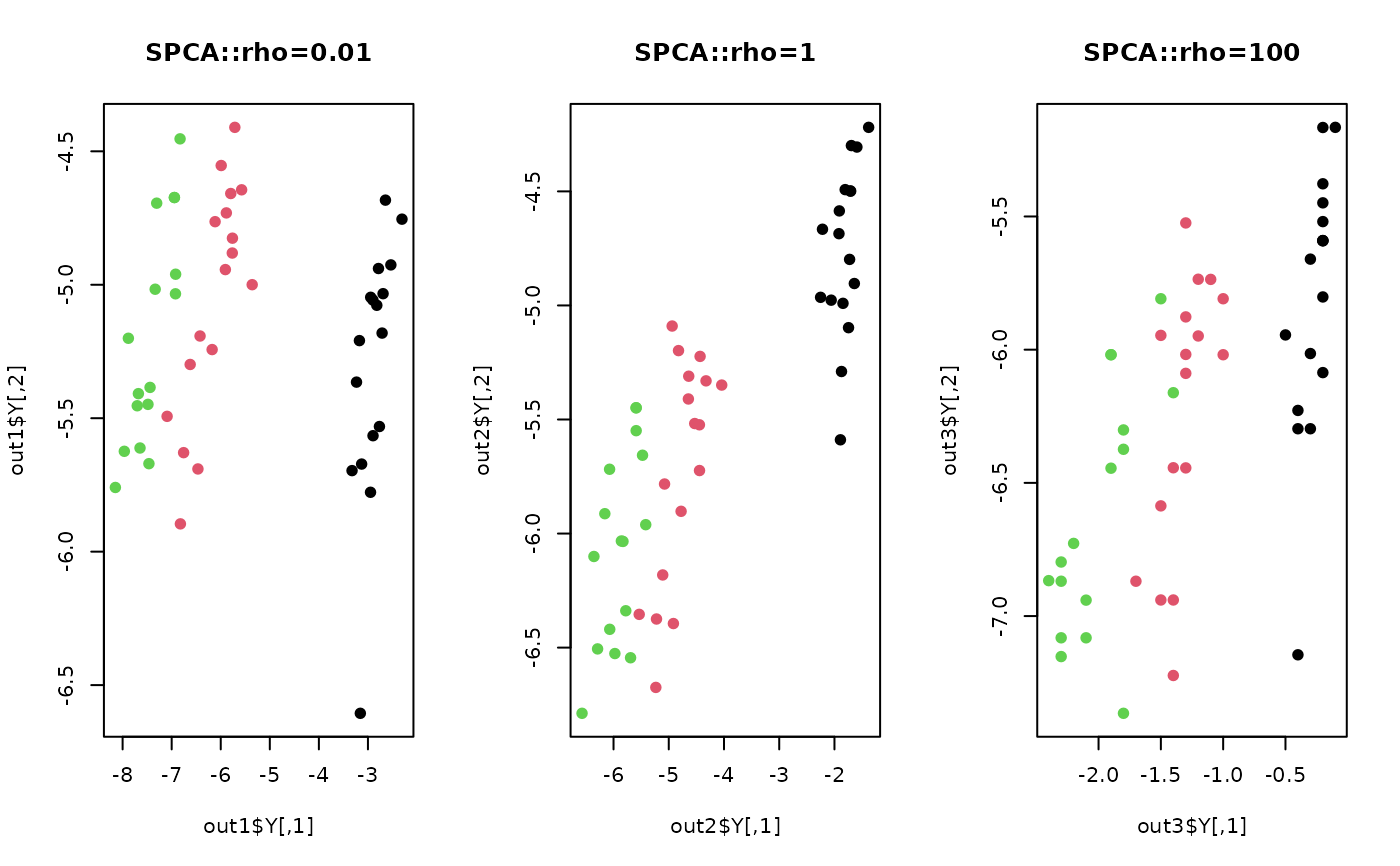 par(opar)
# }
par(opar)
# }