Unlike original principal component analysis (do.pca), this algorithm implements
a supervised version using response information for feature selection. For each feature/column,
its normalized association with response variable is computed and the features with
large magnitude beyond threshold are selected. From the selected submatrix,
regular PCA is applied for dimension reduction.
do.spc(
X,
response,
ndim = 2,
preprocess = c("center", "whiten", "decorrelate"),
threshold = 0.1
)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- response
a length-\(n\) vector of response variable.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is
center. See alsoaux.preprocessfor more details.- threshold
a threshold value to cut off normalized association between covariates and response.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Bair E, Hastie T, Paul D, Tibshirani R (2006). “Prediction by Supervised Principal Components.” Journal of the American Statistical Association, 101(473), 119–137.
Examples
## generate swiss roll with auxiliary dimensions
## it follows reference example from LSIR paper.
set.seed(100)
n = 100
theta = runif(n)
h = runif(n)
t = (1+2*theta)*(3*pi/2)
X = array(0,c(n,10))
X[,1] = t*cos(t)
X[,2] = 21*h
X[,3] = t*sin(t)
X[,4:10] = matrix(runif(7*n), nrow=n)
## corresponding response vector
y = sin(5*pi*theta)+(runif(n)*sqrt(0.1))
## try different threshold values
out1 = do.spc(X, y, threshold=2)
out2 = do.spc(X, y, threshold=5)
out3 = do.spc(X, y, threshold=10)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
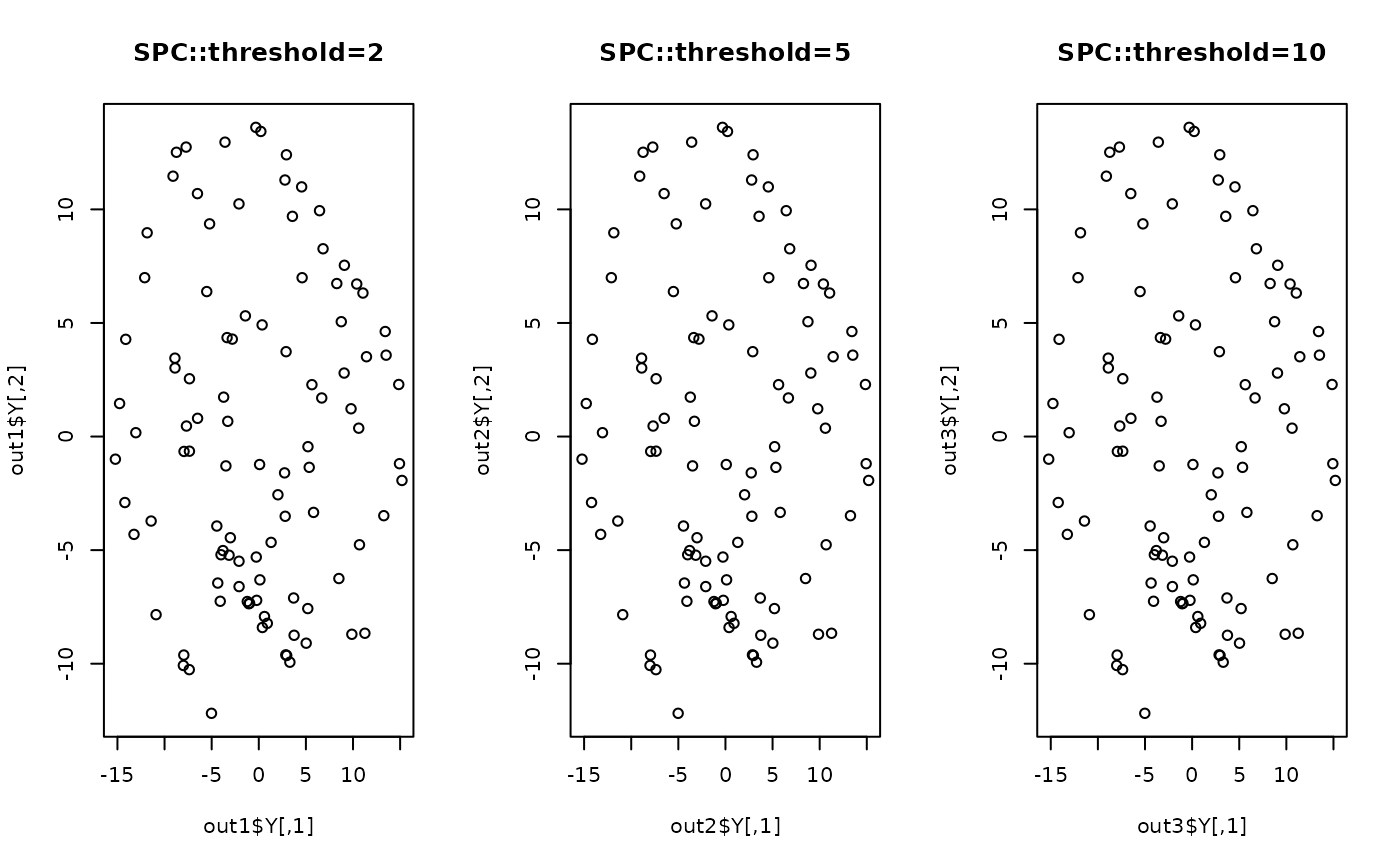
plot(out1$Y, main="SPC::threshold=2")
plot(out2$Y, main="SPC::threshold=5")
plot(out3$Y, main="SPC::threshold=10")
 par(opar)
par(opar)