Given two data sets, Partial Least Squares (PLS) aims at maximizing cross-covariance of latent variables for each data matrix,
therefore it can be considered as supervised methods. As we have two input matrices, do.pls generates two sets of
outputs. Though it is widely used for regression problem, we used it in dimension reduction setting. For
algorithm aspects, we used recursive gram-schmidt orthogonalization in conjunction with extracting projection vectors under
eigen-decomposition formulation, as the problem dimension matters only up to original dimensionality.
For more details, see Wikipedia entry on PLS.
do.pls(data1, data2, ndim = 2)Arguments
Value
a named list containing
- Y1
an \((n\times ndim)\) matrix of projected observations from
data1.- Y2
an \((n\times ndim)\) matrix of projected observations from
data2.- projection1
an \((N\times ndim)\) whose columns are loadings for
data1.- projection2
an \((M\times ndim)\) whose columns are loadings for
data2.- trfinfo1
a list containing information for out-of-sample prediction for
data1.- trfinfo2
a list containing information for out-of-sample prediction for
data2.- eigvals
a vector of eigenvalues for iterative decomposition.
References
Wold H (1975). “Path Models with Latent Variables: The NIPALS Approach.” In Quantitative Sociology, 307–357. Elsevier. ISBN 978-0-12-103950-9.
Rosipal R, Krämer N (2006). “Overview and Recent Advances in Partial Least Squares.” In Saunders C, Grobelnik M, Gunn S, Shawe-Taylor J (eds.), Subspace, Latent Structure and Feature Selection: Statistical and Optimization Perspectives Workshop, SLSFS 2005, Bohinj, Slovenia, February 23-25, 2005, Revised Selected Papers, 34–51. Springer Berlin Heidelberg, Berlin, Heidelberg. ISBN 978-3-540-34138-3.
See also
Examples
## generate 2 normal data matrices
mat1 = matrix(rnorm(100*12),nrow=100)+10 # 12-dim normal
mat2 = matrix(rnorm(100*6), nrow=100)-10 # 6-dim normal
## project onto 2 dimensional space for each data
output = do.pls(mat1, mat2, ndim=2)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,2))
plot(output$Y1, main="proj(mat1)")
plot(output$Y2, main="proj(mat2)")
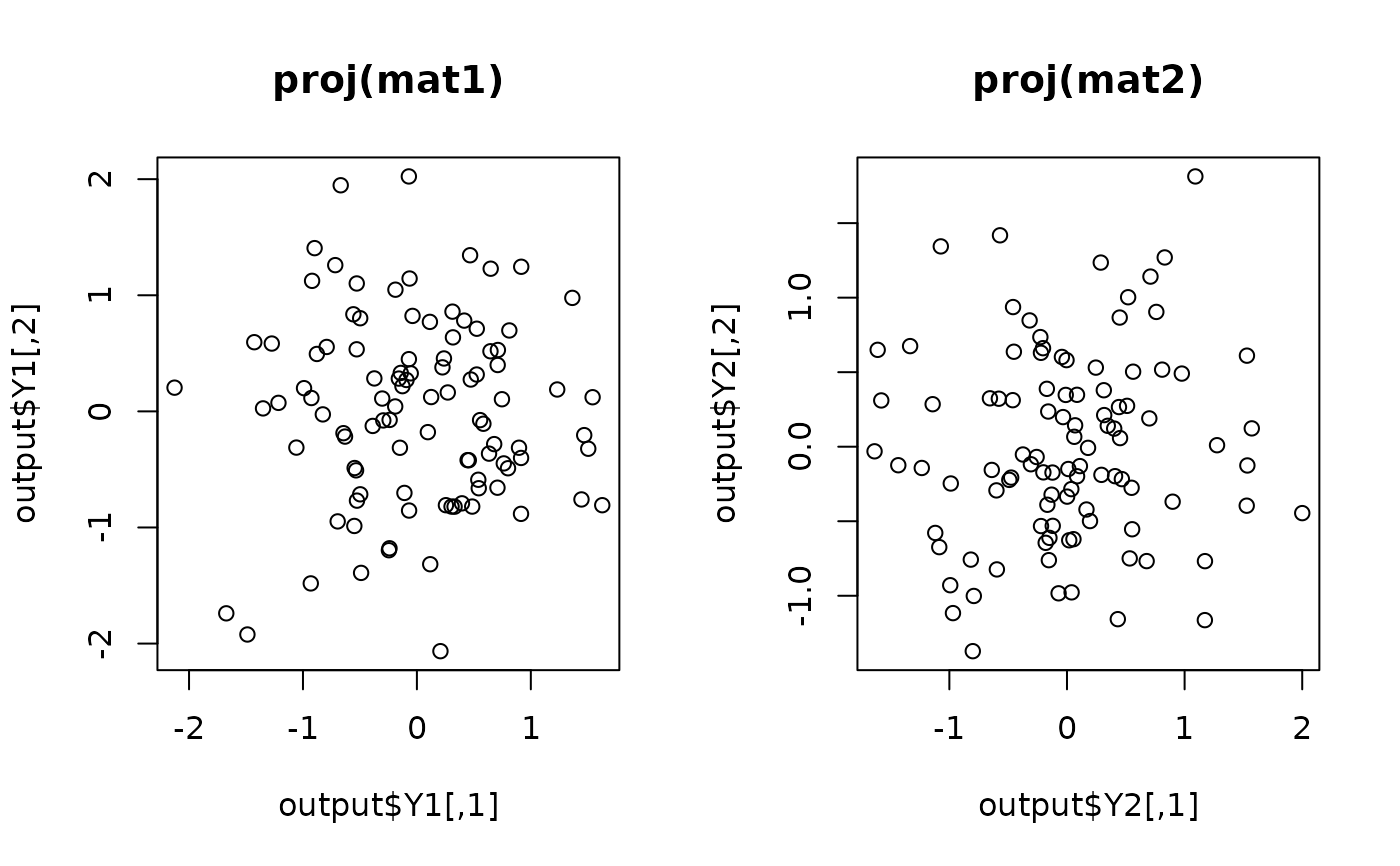 par(opar)
par(opar)