Canonical Correlation Analysis (CCA) is similar to Partial Least Squares (PLS), except for one objective; while PLS focuses on maximizing covariance, CCA maximizes the correlation. This difference sometimes incurs quite distinct results compared to PLS. For algorithm aspects, we used recursive gram-schmidt orthogonalization in conjunction with extracting projection vectors under eigen-decomposition formulation, as the problem dimension matters only up to original dimensionality.
do.cca(data1, data2, ndim = 2)Arguments
Value
a named list containing
- Y1
an \((n\times ndim)\) matrix of projected observations from
data1.- Y2
an \((n\times ndim)\) matrix of projected observations from
data2.- projection1
a \((N\times ndim)\) whose columns are loadings for
data1.- projection2
a \((M\times ndim)\) whose columns are loadings for
data2.- trfinfo1
a list containing information for out-of-sample prediction for
data1.- trfinfo2
a list containing information for out-of-sample prediction for
data2.- eigvals
a vector of eigenvalues for iterative decomposition.
References
Hotelling H (1936). “RELATIONS BETWEEN TWO SETS OF VARIATES.” Biometrika, 28(3-4), 321–377.
See also
Examples
## generate 2 normal data matrices
set.seed(100)
mat1 = matrix(rnorm(100*12),nrow=100)+10 # 12-dim normal
mat2 = matrix(rnorm(100*6), nrow=100)-10 # 6-dim normal
## project onto 2 dimensional space for each data
output = do.cca(mat1, mat2, ndim=2)
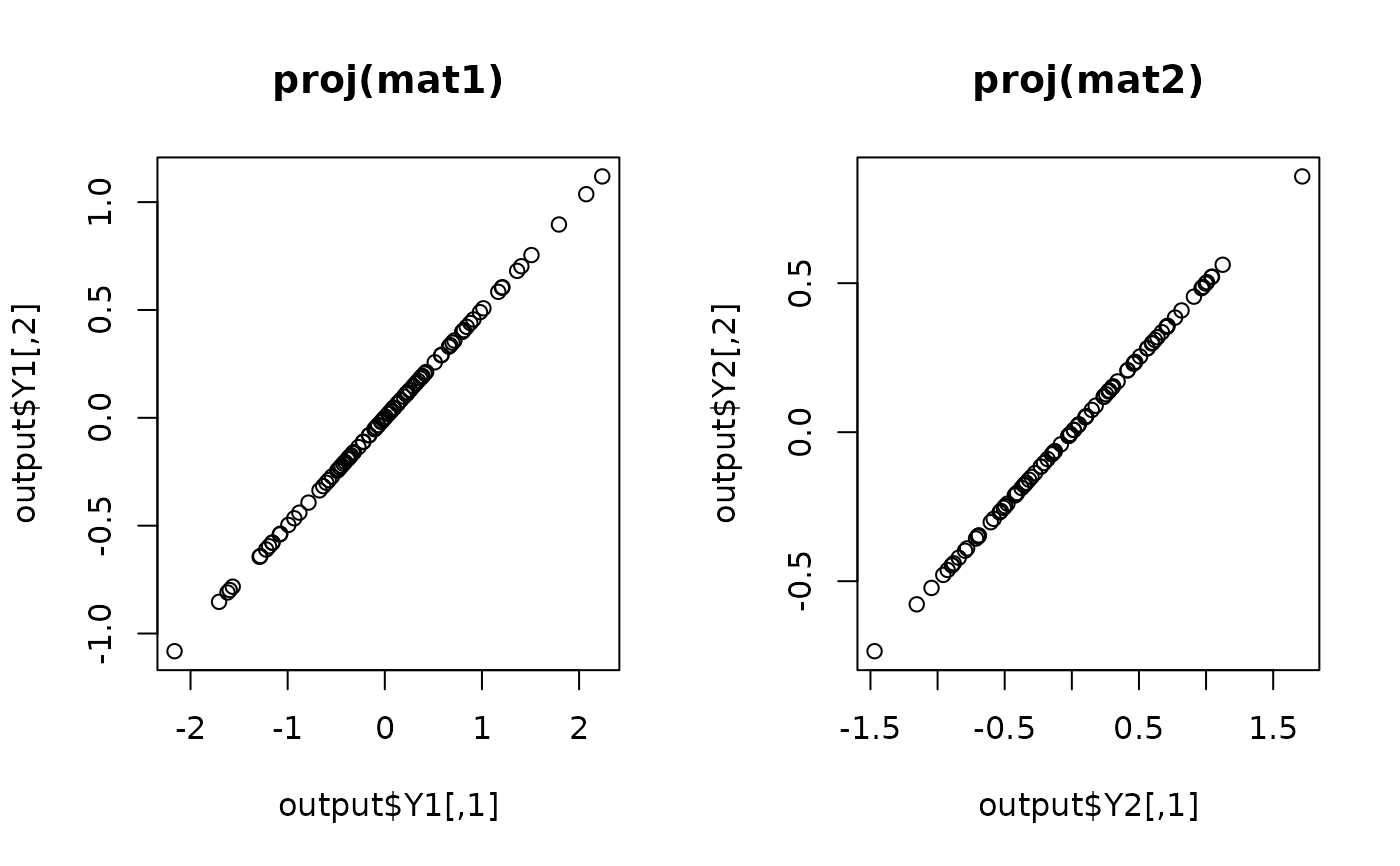
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,2))
plot(output$Y1, main="proj(mat1)")
plot(output$Y2, main="proj(mat2)")
 par(opar)
par(opar)