Principal Component Analysis exploits sample covariance matrix whose
eigenvectors and eigenvalues are principal components and projected
variance, correspondingly. Given varratio, it thresholds the
accumulated variance and selects the estimated dimension. Note that other than
linear submanifold case, the naive selection scheme from this algorithm
lacks flexibility in discovering intrinsic dimension.
est.pcathr(X, varratio = 0.95)Arguments
Value
a named list containing containing
- estdim
estimated dimension according to
varratio.- values
eigenvalues of sample covariance matrix.
See also
Examples
## generate 3-dimensional normal data
X = matrix(rnorm(100*3), nrow=100)
## replicate 3 times with translations
Y = cbind(X-10,X,X+10)
## use PCA thresholding estimation with 95% variance explainability
## desired return is for dimension 3.
output = est.pcathr(Y)
pmessage = paste("* estimated dimension is ",output$estdim, sep="")
print(pmessage)
#> [1] "* estimated dimension is 3"
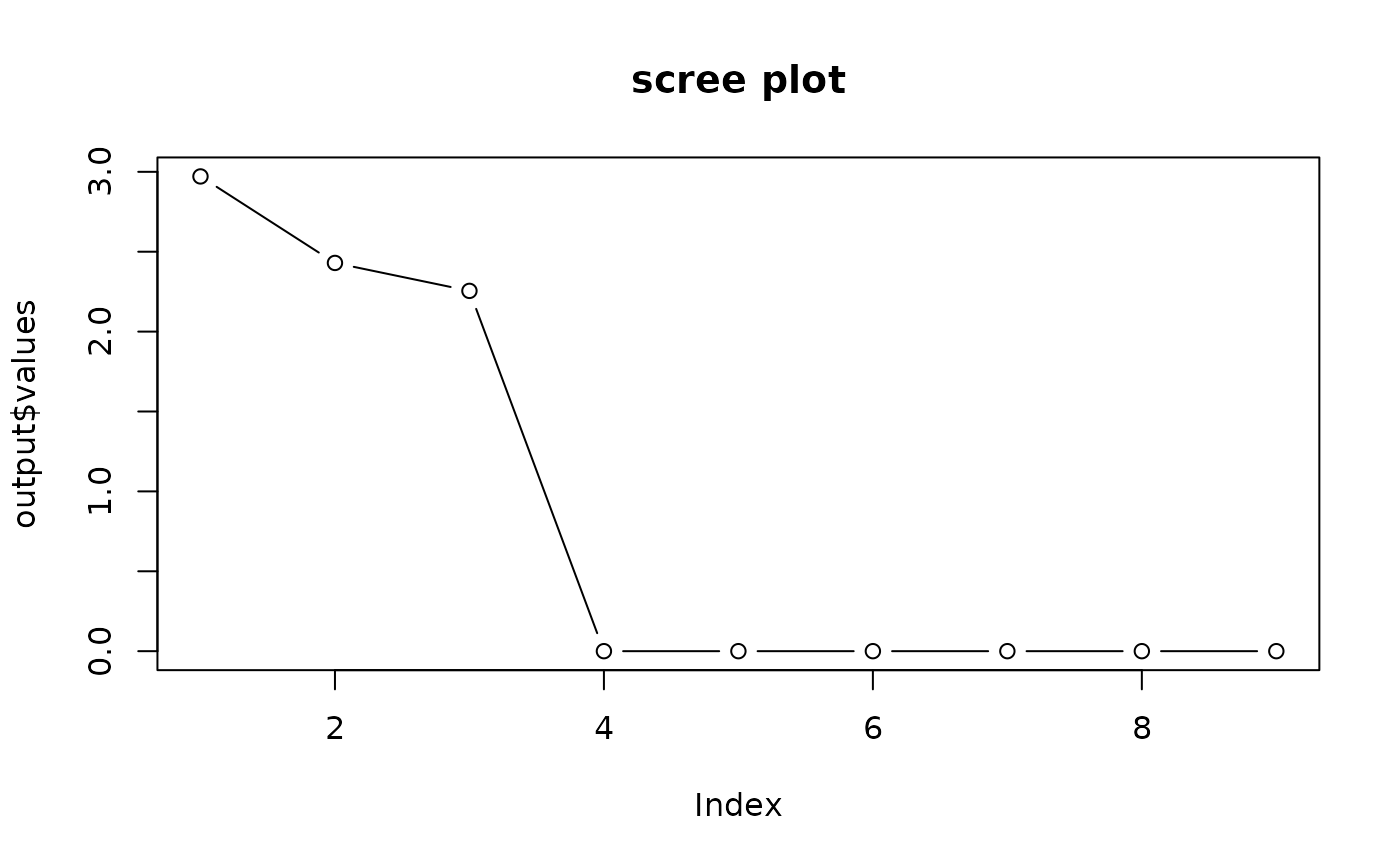
## use screeplot
opar <- par(no.readonly=TRUE)
plot(output$values, main="scree plot", type="b")
 par(opar)
par(opar)