Low-Rank Subspace Clustering (LRSC) constructs the connectivity of the data by solving $$\textrm{min}_C \|C\|_*\quad\textrm{such that}\quad A=AC,~C=C^\top$$ for the uncorrupted data scenario where \(A\) is a column-stacked data matrix. In the current implementation, the first equality constraint for reconstructiveness of the data can be relaxed by solving $$\textrm{min}_C \|C\|_* + \frac{\tau}{2} \|A-AC\|_F^2 \quad\textrm{such that}\quad C=C^\top$$ controlled by the regularization parameter \(\tau\). If you are interested in enabling a more general class of the problem suggested by authors, please contact maintainer of the package.
LRSC(data, k = 2, type = c("relaxed", "exact"), tau = 1)
Arguments
| data | an \((n\times p)\) matrix of row-stacked observations. |
|---|---|
| k | the number of clusters (default: 2). |
| type | type of the problem to be solved. |
| tau | regularization parameter for relaxed-constraint problem. |
Value
a named list of S3 class T4cluster containing
- cluster
a length-\(n\) vector of class labels (from \(1:k\)).
- algorithm
name of the algorithm.
Details
$$\textrm{min}_C \|C\|_*\quad\textrm{such that}\quad D=DC$$ for column-stacked data matrix \(D\) and \(\|\cdot \|_*\) is the nuclear norm which is relaxation of the rank condition. If you are interested in full implementation of the algorithm with sparse outliers and noise, please contact the maintainer.
References
Vidal R, Favaro P (2014). “Low Rank Subspace Clustering (LRSC).” Pattern Recognition Letters, 43, 47--61. ISSN 01678655.
Examples
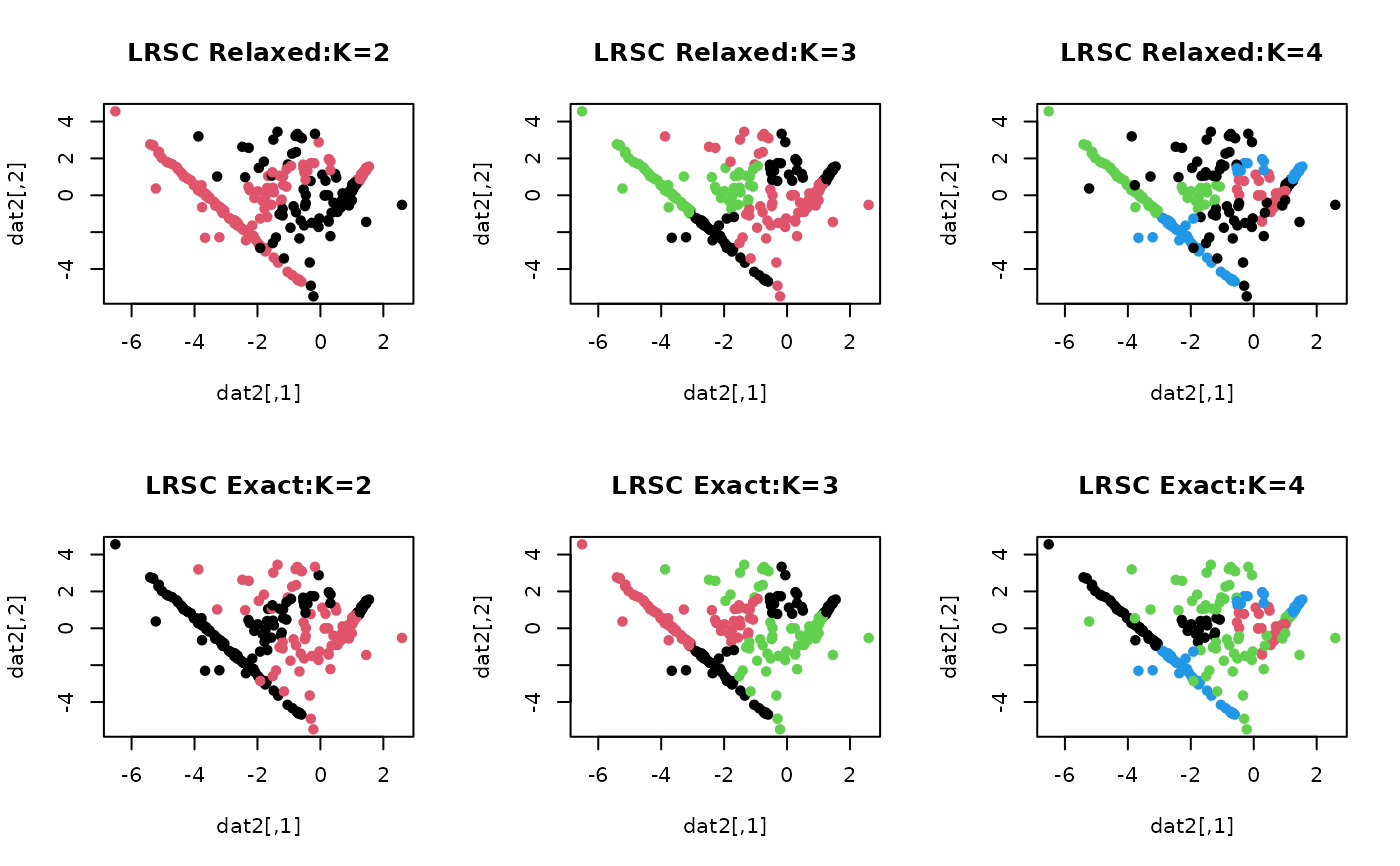
# \donttest{ ## generate a toy example set.seed(10) tester = genLP(n=100, nl=2, np=1, iso.var=0.1) data = tester$data label = tester$class ## do PCA for data reduction proj = base::eigen(stats::cov(data))$vectors[,1:2] dat2 = data%*%proj ## run LRSC algorithm with k=2,3,4 with relaxed/exact solvers out2rel = LRSC(data, k=2, type="relaxed") out3rel = LRSC(data, k=3, type="relaxed") out4rel = LRSC(data, k=4, type="relaxed") out2exc = LRSC(data, k=2, type="exact") out3exc = LRSC(data, k=3, type="exact") out4exc = LRSC(data, k=4, type="exact") ## extract label information lab2rel = out2rel$cluster lab3rel = out3rel$cluster lab4rel = out4rel$cluster lab2exc = out2exc$cluster lab3exc = out3exc$cluster lab4exc = out4exc$cluster ## visualize opar <- par(no.readonly=TRUE) par(mfrow=c(2,3)) plot(dat2, pch=19, cex=0.9, col=lab2rel, main="LRSC Relaxed:K=2") plot(dat2, pch=19, cex=0.9, col=lab3rel, main="LRSC Relaxed:K=3") plot(dat2, pch=19, cex=0.9, col=lab4rel, main="LRSC Relaxed:K=4") plot(dat2, pch=19, cex=0.9, col=lab2exc, main="LRSC Exact:K=2") plot(dat2, pch=19, cex=0.9, col=lab3exc, main="LRSC Exact:K=3") plot(dat2, pch=19, cex=0.9, col=lab4exc, main="LRSC Exact:K=4")