Local Tangent Space Alignment, or LTSA in short, is a nonlinear dimensionality reduction method that mimicks the behavior of low-dimensional manifold embedded in high-dimensional space. Similar to LLE, LTSA computes tangent space using nearest neighbors of a given data point, and a multiple of tangent spaces are gathered to to find an embedding that aligns the tangent spaces in target dimensional space.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- symmetric
one of
"intersect","union"or"asymmetric"is supported. Default is"union". See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- eigvals
a vector of eigenvalues from the final decomposition.
References
Zhang T, Yang J, Zhao D, Ge X (2007). “Linear Local Tangent Space Alignment and Application to Face Recognition.” Neurocomputing, 70(7-9), 1547–1553.
Examples
# \donttest{
## generate data
set.seed(100)
X <- aux.gensamples(dname="cswiss",n=100)
## 1. use 10%-connected graph
output1 <- do.ltsa(X,ndim=2)
## 2. use 25%-connected graph
output2 <- do.ltsa(X,ndim=2,type=c("proportion",0.25))
## 3. use 50%-connected graph
output3 <- do.ltsa(X,ndim=2,type=c("proportion",0.50))
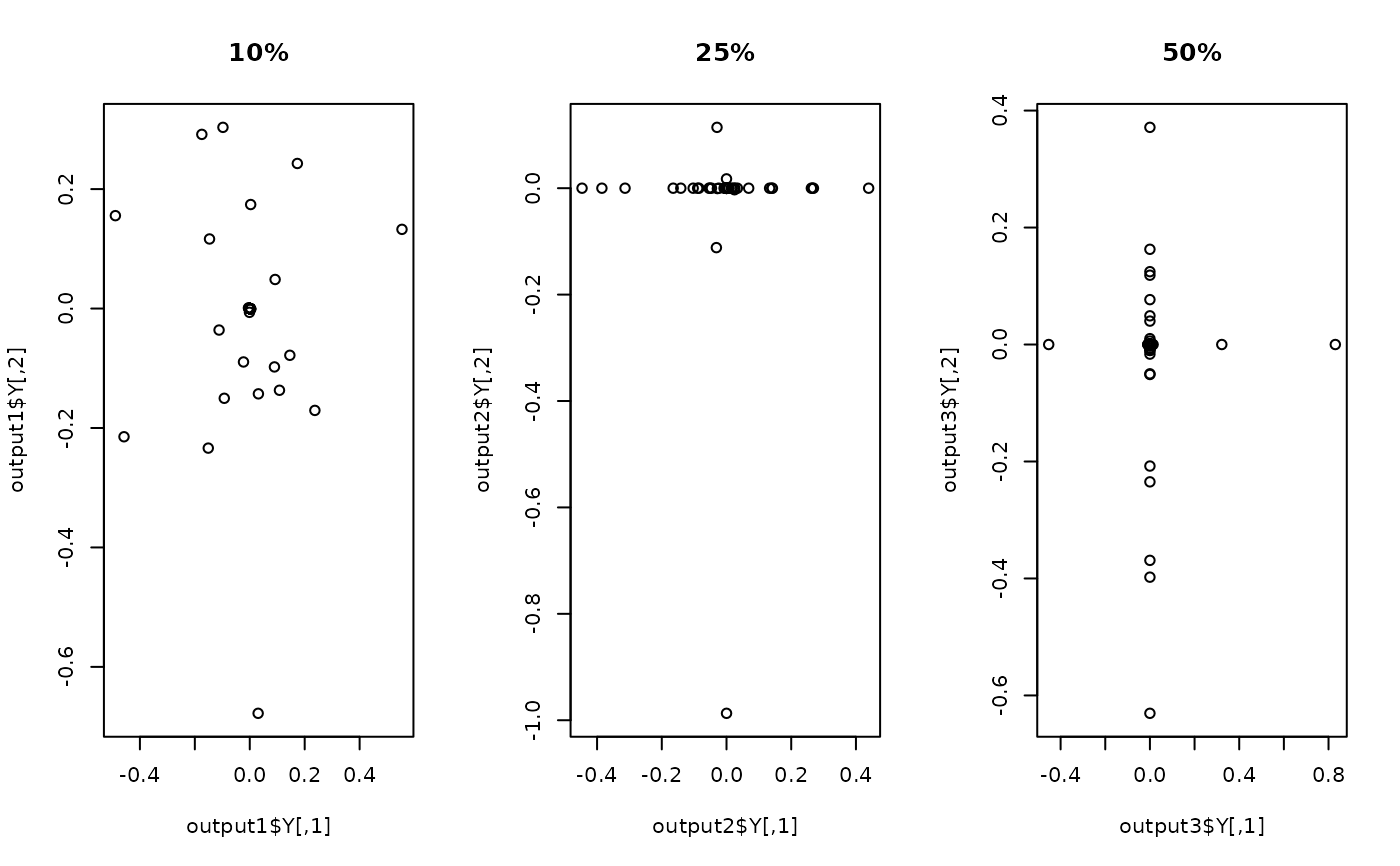
## Visualize three different projections
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
plot(output1$Y, main="10%")
plot(output2$Y, main="25%")
plot(output3$Y, main="50%")
 par(opar)
# }
par(opar)
# }