Kernel LFDA is a nonlinear extension of LFDA method using kernel trick. It applies conventional kernel method
to extend excavation of hidden patterns in a more flexible manner in tradeoff of computational load. For simplicity,
only the gaussian kernel parametrized by its bandwidth t is supported.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- label
a length-\(n\) vector of data class labels.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- symmetric
one of
"intersect","union"or"asymmetric"is supported. Default is"union". See alsoaux.graphnbdfor more details.- localscaling
TRUEto use local scaling method for construction affinity matrix,FALSEfor binary affinity.- t
bandwidth parameter for heat kernel in \((0,\infty)\).
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
References
Sugiyama M (2006). “Local Fisher Discriminant Analysis for Supervised Dimensionality Reduction.” In Proceedings of the 23rd International Conference on Machine Learning, 905–912.
Zelnik-manor L, Perona P (2005). “Self-Tuning Spectral Clustering.” In Saul LK, Weiss Y, Bottou L (eds.), Advances in Neural Information Processing Systems 17, 1601–1608. MIT Press.
See also
Examples
# \donttest{
## generate 3 different groups of data X and label vector
set.seed(100)
x1 = matrix(rnorm(4*10), nrow=10)-20
x2 = matrix(rnorm(4*10), nrow=10)
x3 = matrix(rnorm(4*10), nrow=10)+20
X = rbind(x1, x2, x3)
label = rep(1:3, each=10)
## try different affinity matrices
out1 = do.klfda(X, label, t=0.1)
out2 = do.klfda(X, label, t=1)
out3 = do.klfda(X, label, t=10)
## visualize
opar = par(no.readonly=TRUE)
par(mfrow=c(1,3))
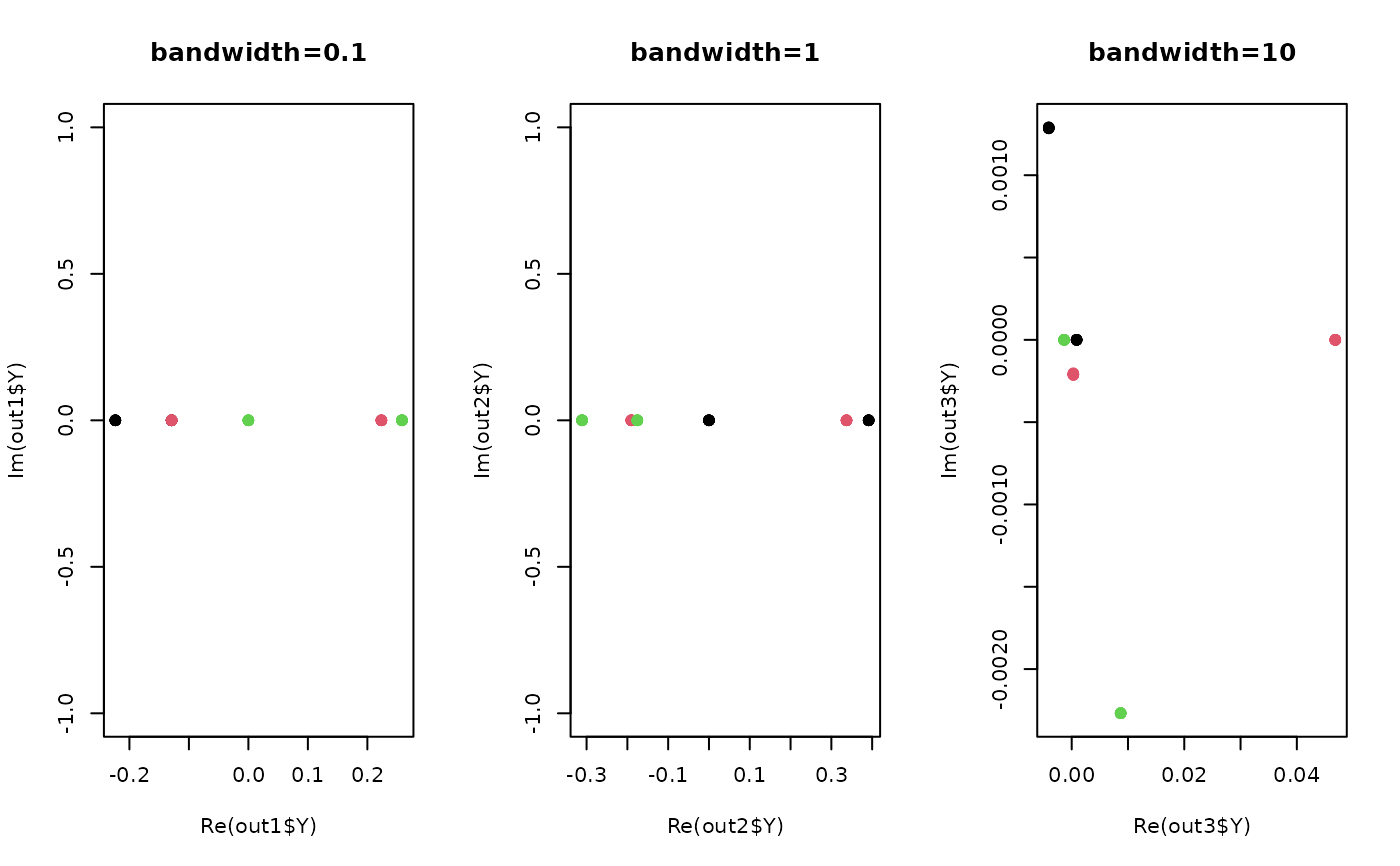
plot(out1$Y, pch=19, col=label, main="bandwidth=0.1")
plot(out2$Y, pch=19, col=label, main="bandwidth=1")
plot(out3$Y, pch=19, col=label, main="bandwidth=10")
 par(opar)
# }
par(opar)
# }