Kernel Entropy Component Analysis(KECA) is a kernel method of dimensionality reduction.
Unlike Kernel PCA(do.kpca), it utilizes eigenbasis of kernel matrix \(K\)
in accordance with indices of largest Renyi quadratic entropy in which entropy for
\(j\)-th eigenpair is defined to be \(\sqrt{\lambda_j}e_j^T 1_n\), where \(e_j\) is
\(j\)-th eigenvector of an uncentered kernel matrix \(K\).
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- kernel
a vector containing name of a kernel and corresponding parameters. See also
aux.kernelcovfor complete description of Kernel Trick.- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- entropy
a length-
ndimvector of estimated entropy values.
References
Jenssen R (2010). “Kernel Entropy Component Analysis.” IEEE Transactions on Pattern Analysis and Machine Intelligence, 32(5), 847–860.
See also
Examples
# \donttest{
## load iris data
data(iris)
set.seed(100)
subid = sample(1:150,50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
## 1. standard KECA with gaussian kernel
output1 <- do.keca(X,ndim=2)
## 2. gaussian kernel with large bandwidth
output2 <- do.keca(X,ndim=2,kernel=c("gaussian",5))
## 3. use laplacian kernel
output3 <- do.keca(X,ndim=2,kernel=c("laplacian",1))
## Visualize three different projections
opar = par(no.readonly=TRUE)
par(mfrow=c(1,3))
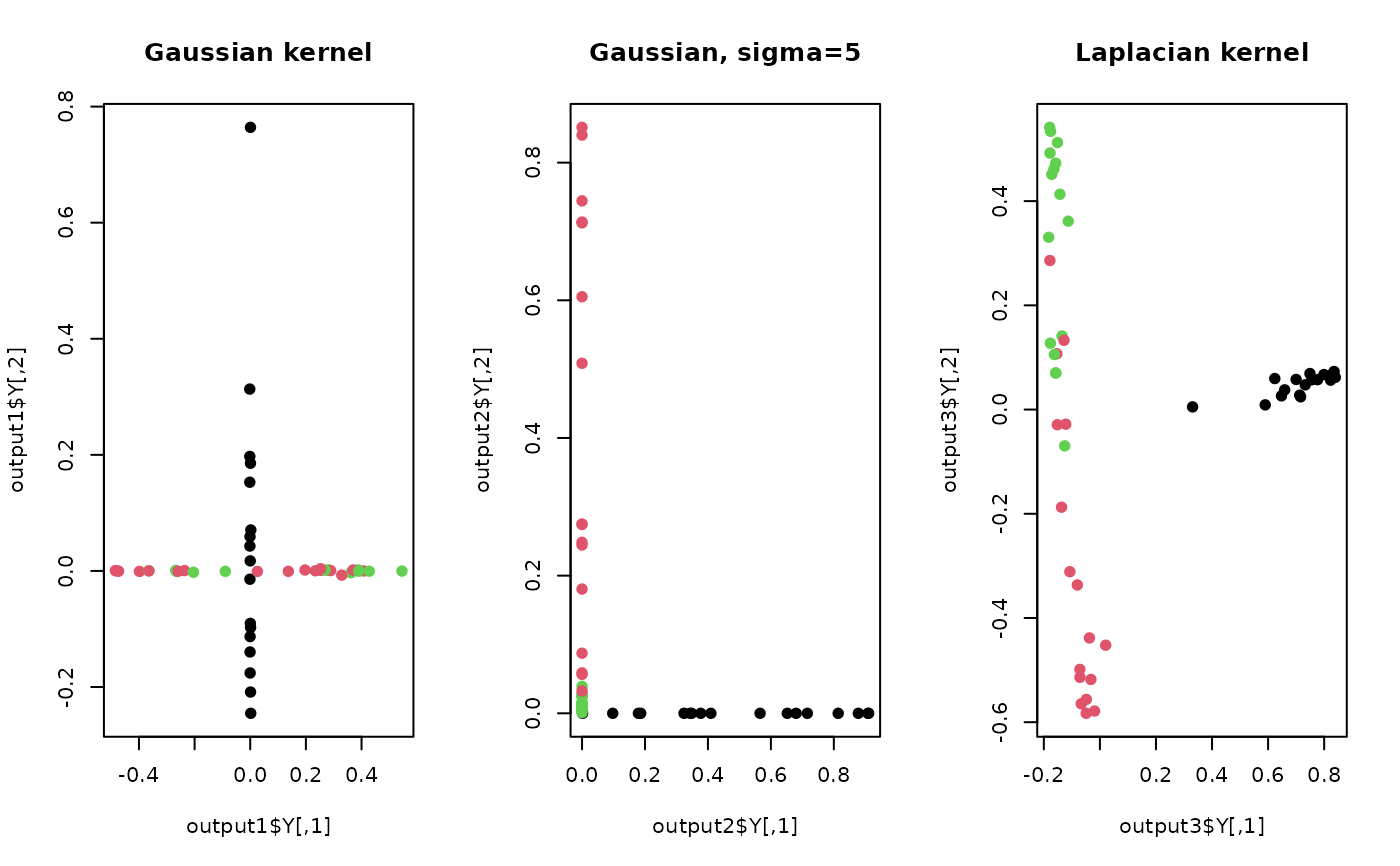
plot(output1$Y, pch=19, col=label, main="Gaussian kernel")
plot(output2$Y, pch=19, col=label, main="Gaussian, sigma=5")
plot(output3$Y, pch=19, col=label, main="Laplacian kernel")
 par(opar)
# }
par(opar)
# }