Hyperbolic Distance Recovery and Approximation, also known as hydra in short,
implements embedding of distance-based data into hyperbolic space represented as the Poincare disk,
which is interior of a hypersphere.
do.hydra(X, ndim = 2, ...)Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension (default: 2).
- ...
extra parameters including
- kappa
embedding curvature, which is a nonnegative number (default: 1).
- iso.adjust
perform isotropic adjustment. If
ndim=2, default isFALSE. Otherwise,TRUEis used as default.
Value
a named Rdimtools S3 object containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations in the Poincare disk.
- algorithm
name of the algorithm.
References
Keller-Ressel M, Nargang S (2020). “Hydra: A Method for Strain-Minimizing Hyperbolic Embedding of Network- and Distance-Based Data.” Journal of Complex Networks, 8(1), cnaa002. ISSN 2051-1329.
Examples
# \donttest{
## load iris data
data(iris)
X = as.matrix(iris[,1:4])
lab = as.factor(iris[,5])
## multiple runs with varying curvatures
embed1 <- do.hydra(X, kappa=0.1)
embed2 <- do.hydra(X, kappa=1)
embed3 <- do.hydra(X, kappa=10)
## Visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3), pty="s")
plot(embed1$Y , col=lab, pch=19, main="kappa=0.1")
plot(embed2$Y , col=lab, pch=19, main="kappa=1")
plot(embed3$Y , col=lab, pch=19, main="kappa=10")
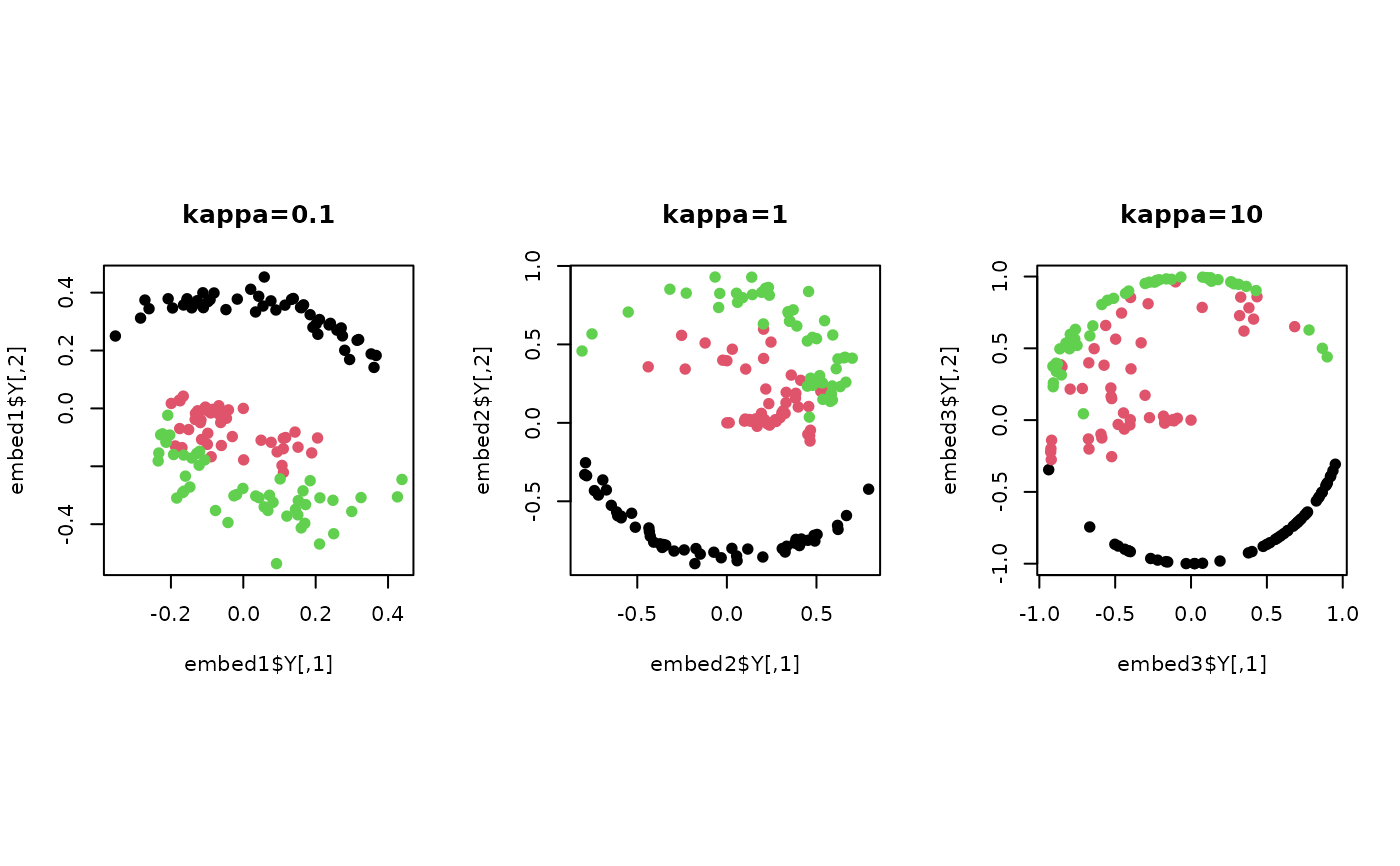 par(opar)
# }
par(opar)
# }