Distinguishing Variance Embedding (DVE) is an unsupervised nonlinear manifold learning method. It can be considered as a balancing method between Maximum Variance Unfolding and Laplacian Eigenmaps. The algorithm unfolds the data by maximizing the global variance subject to the locality-preserving constraint. Instead of defining certain kernel, it applies local scaling scheme in that it automatically computes adaptive neighborhood-based kernel bandwidth.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
References
Wang Q, Li J (2009). “Combining Local and Global Information for Nonlinear Dimensionality Reduction.” Neurocomputing, 72(10-12), 2235–2241.
Qinggang W, Jianwei L, Xuchu W (2010). “Distinguishing Variance Embedding.” Image and Vision Computing, 28(6), 872–880.
Examples
# \donttest{
## generate swiss-roll dataset of size 100
set.seed(100)
X <- aux.gensamples(dname="crown", n=100)
## try different nbd size
out1 <- do.dve(X, type=c("proportion",0.5))
out2 <- do.dve(X, type=c("proportion",0.7))
out3 <- do.dve(X, type=c("proportion",0.9))
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
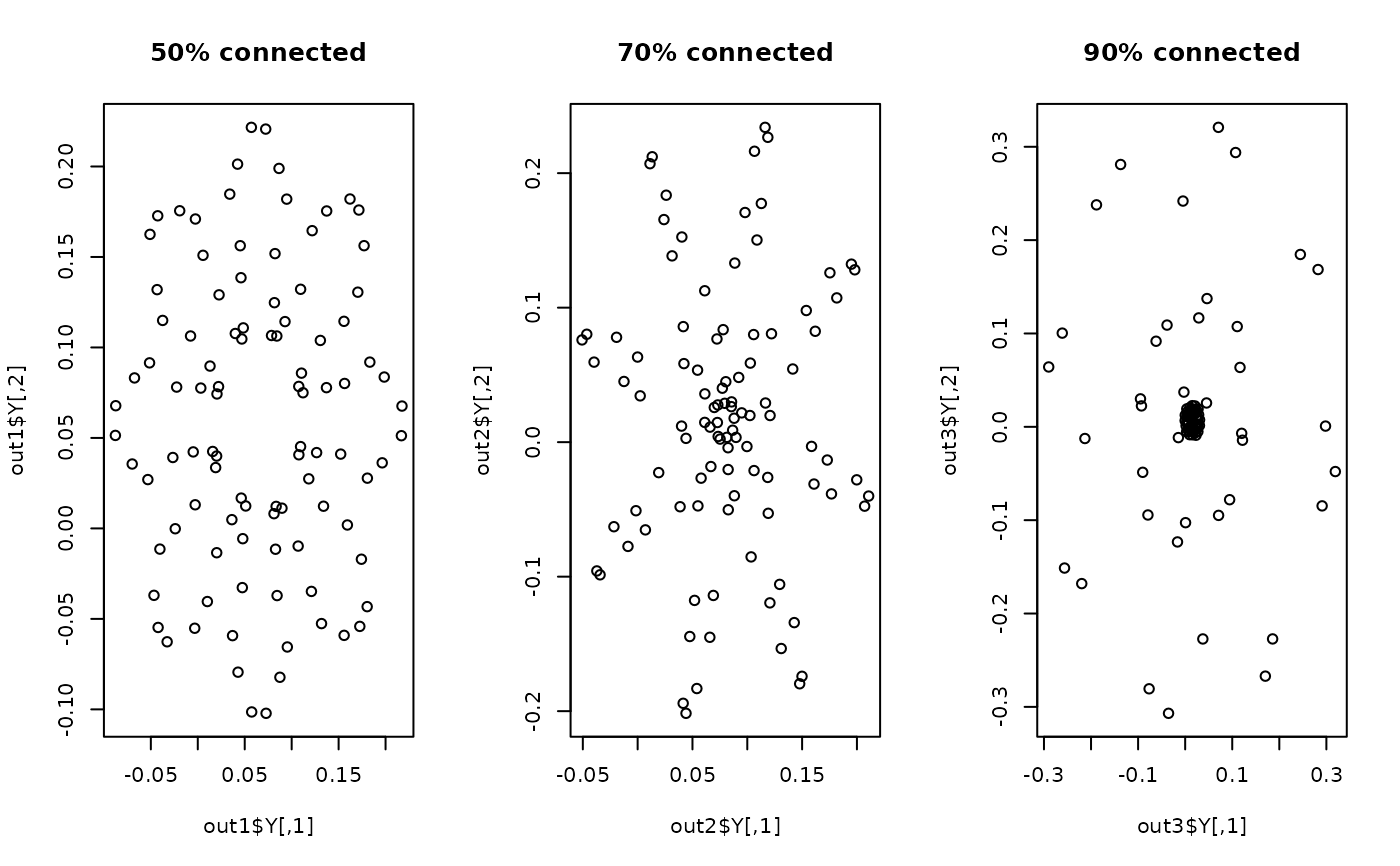
plot(out1$Y, main="50% connected")
plot(out2$Y, main="70% connected")
plot(out3$Y, main="90% connected")
 par(opar)
# }
par(opar)
# }