Unsupervised Discriminant Projection (UDP) aims finding projection that balances local and global scatter. Even though the name contains the word Discriminant, this algorithm is unsupervised. The term there reflects its algorithmic tactic to discriminate distance points not in the neighborhood of each data point. It performs PCA as intermittent preprocessing for rank singularity issue. Authors clearly mentioned that it is inspired by Locality Preserving Projection, which minimizes the local scatter only.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
- interimdim
the number of PCA target dimension used in preprocessing.
References
Yang J, Zhang D, Yang J, Niu B (2007). “Globally Maximizing, Locally Minimizing: Unsupervised Discriminant Projection with Applications to Face and Palm Biometrics.” IEEE Transactions on Pattern Analysis and Machine Intelligence, 29(4), 650–664.
See also
Examples
## load iris data
data(iris)
set.seed(100)
subid = sample(1:150,50)
X = as.matrix(iris[subid,1:4])
label = as.factor(iris[subid,5])
## use different connectivity level
out1 <- do.udp(X, type=c("proportion",0.05))
out2 <- do.udp(X, type=c("proportion",0.10))
out3 <- do.udp(X, type=c("proportion",0.25))
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
plot(out1$Y, col=label, pch=19, main="connectivity 5%")
plot(out2$Y, col=label, pch=19, main="connectivity 10%")
plot(out3$Y, col=label, pch=19, main="connectivity 25%")
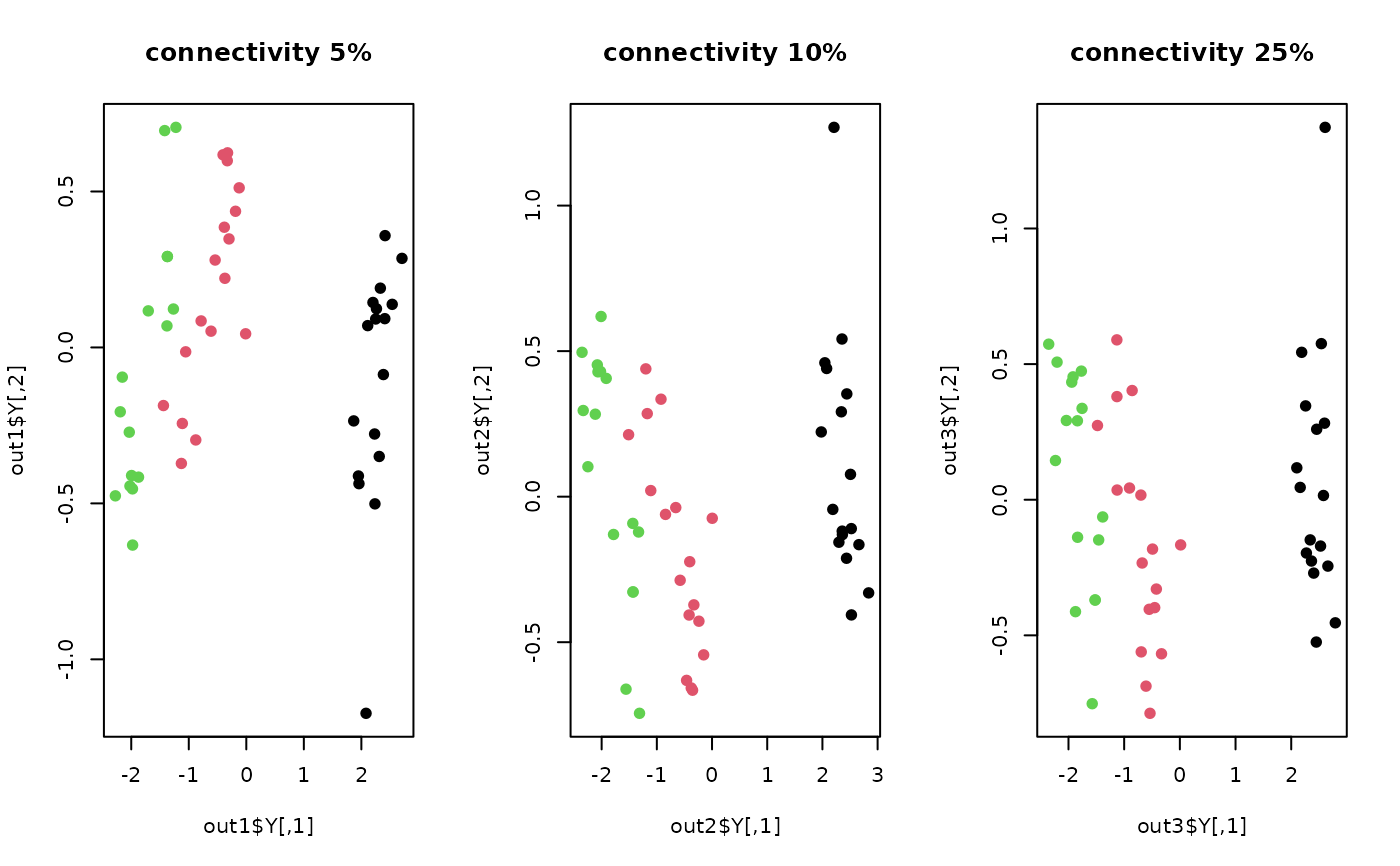 par(opar)
par(opar)