Sliced Average Variance Estimation (SAVE) is a supervised linear dimension reduction method. It is based on sufficiency principle with respect to central subspace concept under the linerity and constant covariance conditions. For more details, see the reference paper.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- response
a length-\(n\) vector of response variable.
- ndim
an integer-valued target dimension.
- h
the number of slices to divide the range of response vector.
- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Dennis Cook R (2000). “Save: A Method for Dimension Reduction and Graphics in Regression.” Communications in Statistics - Theory and Methods, 29(9-10), 2109–2121.
See also
Examples
## generate swiss roll with auxiliary dimensions
## it follows reference example from LSIR paper.
set.seed(100)
n = 50
theta = runif(n)
h = runif(n)
t = (1+2*theta)*(3*pi/2)
X = array(0,c(n,10))
X[,1] = t*cos(t)
X[,2] = 21*h
X[,3] = t*sin(t)
X[,4:10] = matrix(runif(7*n), nrow=n)
## corresponding response vector
y = sin(5*pi*theta)+(runif(n)*sqrt(0.1))
## try with different numbers of slices
out1 = do.save(X, y, h=2)
out2 = do.save(X, y, h=5)
out3 = do.save(X, y, h=10)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
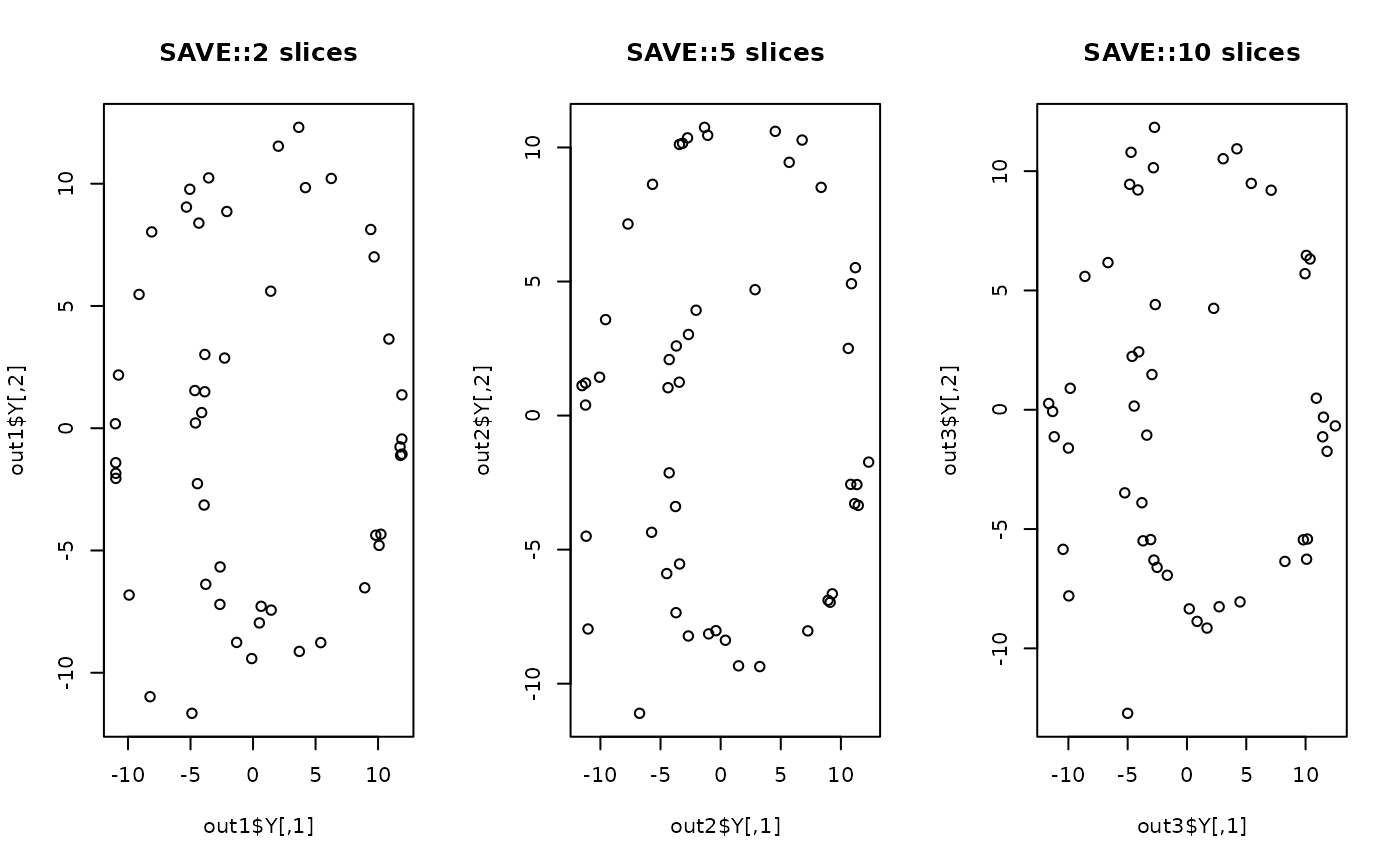
plot(out1$Y, main="SAVE::2 slices")
plot(out2$Y, main="SAVE::5 slices")
plot(out3$Y, main="SAVE::10 slices")
 par(opar)
par(opar)