Modified Orthogonal Discriminant Projection (MODP) is a variant of Orthogonal Discriminant Projection (ODP). Authors argue the assumption in modeling ODP's mechanism to reflect distance and class labeling seem unsound. They propose a modified method to explore the intrinsic structure of original data and enhance the classification ability.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- label
a length-\(n\) vector of data class labels.
- ndim
an integer-valued target dimension.
- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- symmetric
one of
"intersect","union"or"asymmetric"is supported. Default is"union". See alsoaux.graphnbdfor more details.- alpha
balancing parameter of non-local and local scatter in \([0,1]\).
- beta
scaling control parameter for distant pairs of data in \((0,\infty)\).
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
- trfinfo
a list containing information for out-of-sample prediction.
References
Zhang S, Lei Y, Wu Y, Yang J (2011). “Modified Orthogonal Discriminant Projection for Classification.” Neurocomputing, 74(17), 3690–3694.
Examples
## generate 3 different groups of data X and label vector
x1 = matrix(rnorm(4*10), nrow=10)-20
x2 = matrix(rnorm(4*10), nrow=10)
x3 = matrix(rnorm(4*10), nrow=10)+20
X = rbind(x1, x2, x3)
label = rep(1:3, each=10)
## try different beta (scaling control) parameter
out1 = do.modp(X, label, beta=1)
out2 = do.modp(X, label, beta=10)
out3 = do.modp(X, label, beta=100)
## visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
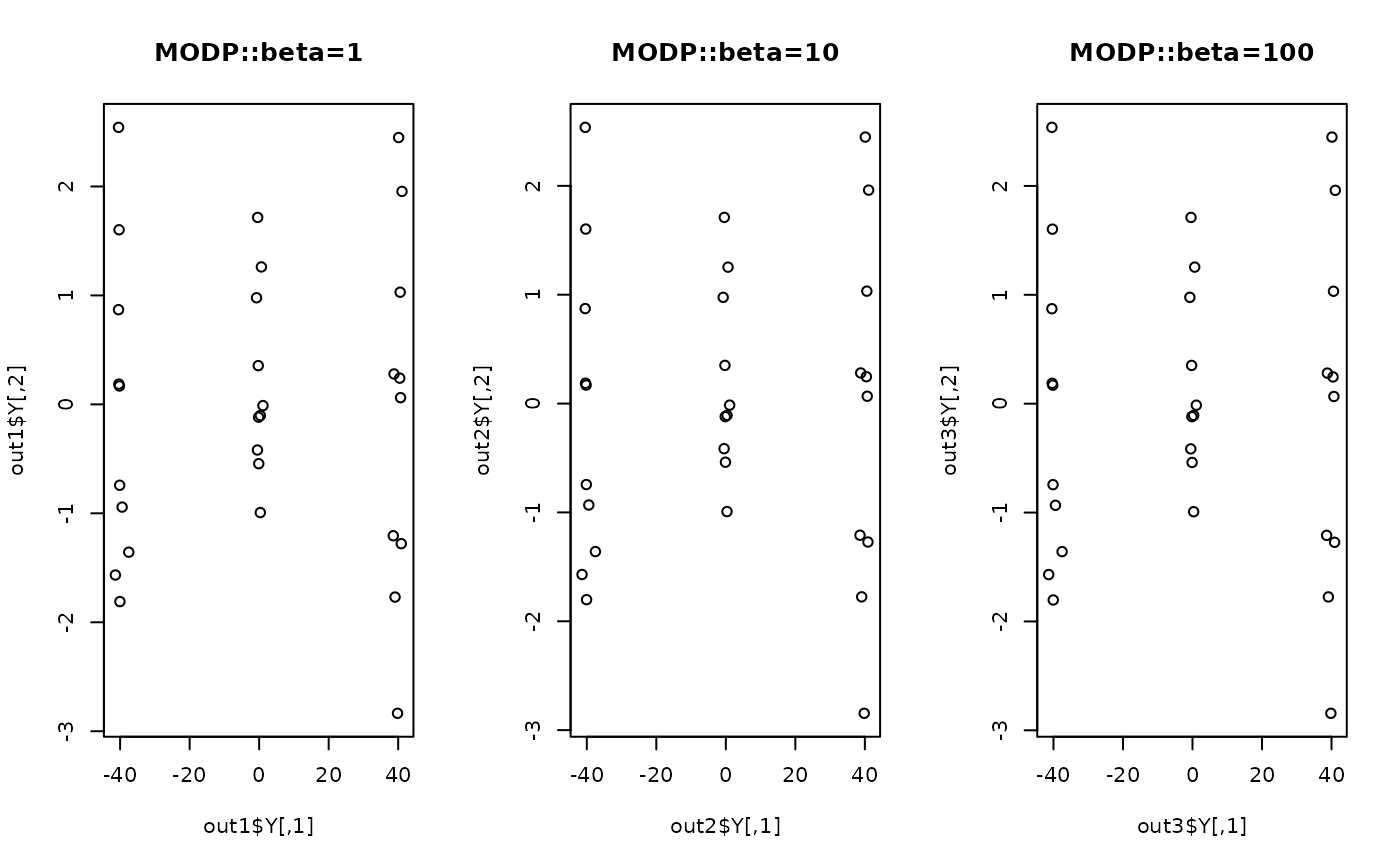
plot(out1$Y, main="MODP::beta=1")
plot(out2$Y, main="MODP::beta=10")
plot(out3$Y, main="MODP::beta=100")
 par(opar)
par(opar)