Locality-Preserved Maximum Information Projection (LPMIP) is an unsupervised linear dimension reduction method
to identify the underlying manifold structure by learning both the within- and between-locality information. The
parameter alpha is balancing the tradeoff between two and the flexibility of this model enables an interpretation
of it as a generalized extension of LPP.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations and columns represent independent variables.
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "null". See also
aux.preprocessfor more details.- sigma
bandwidth parameter for heat kernel in \((0,\infty)\).
- alpha
balancing parameter between two locality information in \([0,1]\).
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Haixian Wang, Sibao Chen, Zilan Hu, Wenming Zheng (2008). “Locality-Preserved Maximum Information Projection.” IEEE Transactions on Neural Networks, 19(4), 571–585.
Examples
## use iris dataset
data(iris)
set.seed(100)
subid <- sample(1:150, 50)
X <- as.matrix(iris[subid,1:4])
lab <- as.factor(iris[subid,5])
## try different neighborhood size
out1 <- do.lpmip(X, ndim=2, type=c("proportion",0.10))
out2 <- do.lpmip(X, ndim=2, type=c("proportion",0.25))
out3 <- do.lpmip(X, ndim=2, type=c("proportion",0.50))
## Visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
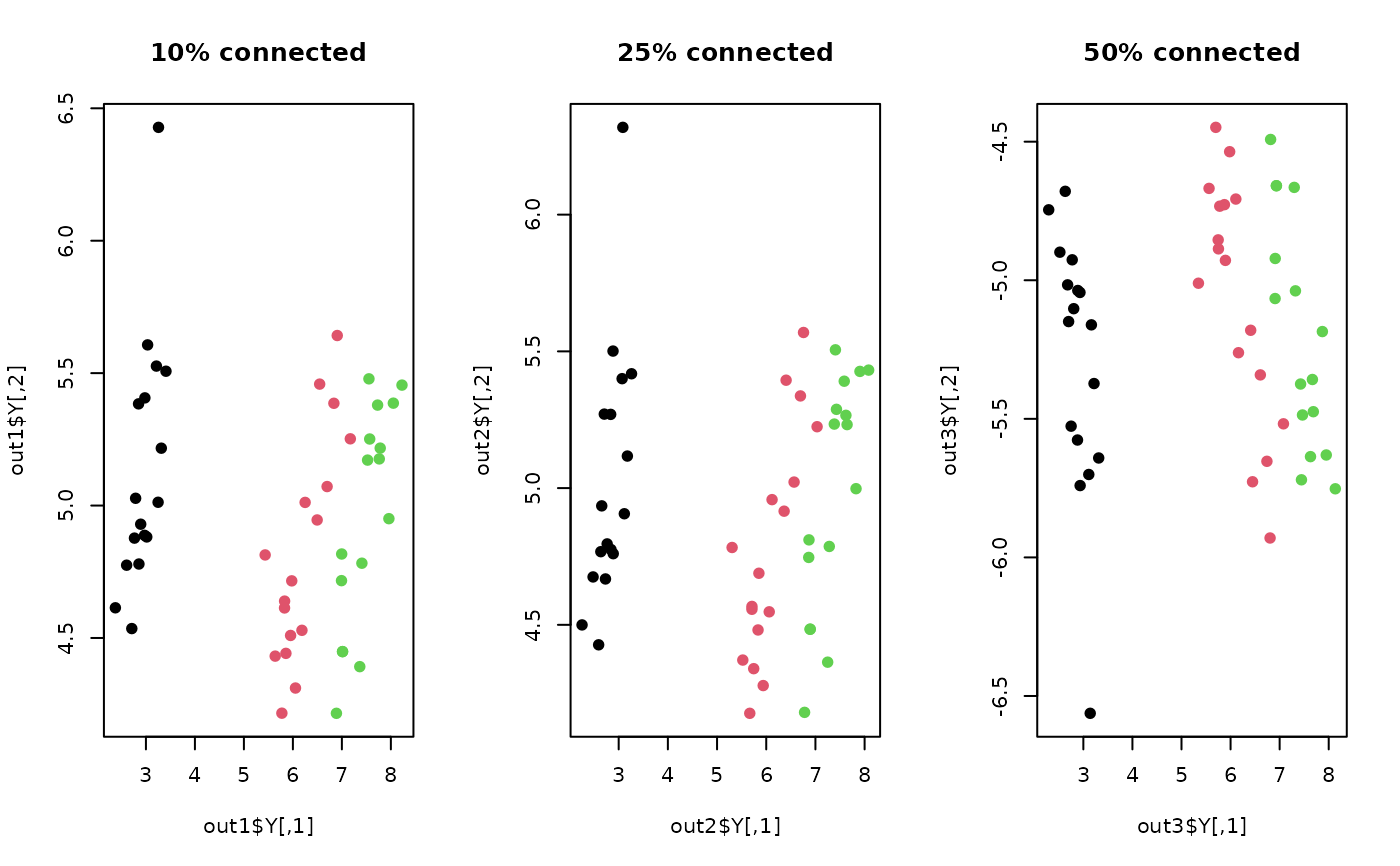
plot(out1$Y, pch=19, col=lab, main="10% connected")
plot(out2$Y, pch=19, col=lab, main="25% connected")
plot(out3$Y, pch=19, col=lab, main="50% connected")
 par(opar)
par(opar)