While Principal Component Analysis (PCA) aims at minimizing global estimation error, Local Learning
Projection (LLP) approach tries to find the projection with the minimal local
estimation error in the sense that each projected datum can be well represented
based on ones neighbors. For the kernel part, we only enabled to use
a gaussian kernel as suggested from the original paper. The parameter lambda
controls possible rank-deficiency of kernel matrix.
Arguments
- X
an \((n\times p)\) matrix or data frame whose rows are observations
- ndim
an integer-valued target dimension.
- type
a vector of neighborhood graph construction. Following types are supported;
c("knn",k),c("enn",radius), andc("proportion",ratio). Default isc("proportion",0.1), connecting about 1/10 of nearest data points among all data points. See alsoaux.graphnbdfor more details.- symmetric
one of
"intersect","union"or"asymmetric"is supported. Default is"union". See alsoaux.graphnbdfor more details.- preprocess
an additional option for preprocessing the data. Default is "center". See also
aux.preprocessfor more details.- t
bandwidth for heat kernel in \((0,\infty)\).
- lambda
regularization parameter for kernel matrix in \([0,\infty)\).
Value
a named list containing
- Y
an \((n\times ndim)\) matrix whose rows are embedded observations.
- trfinfo
a list containing information for out-of-sample prediction.
- projection
a \((p\times ndim)\) whose columns are basis for projection.
References
Wu M, Yu K, Yu S, Schölkopf B (2007). “Local Learning Projections.” In Proceedings of the 24th International Conference on Machine Learning, 1039–1046.
Examples
# \donttest{
## generate data
set.seed(100)
X <- aux.gensamples(n=100, dname="crown")
## test different lambda - regularization - values
out1 <- do.llp(X,ndim=2,lambda=0.1)
out2 <- do.llp(X,ndim=2,lambda=1)
out3 <- do.llp(X,ndim=2,lambda=10)
# visualize
opar <- par(no.readonly=TRUE)
par(mfrow=c(1,3))
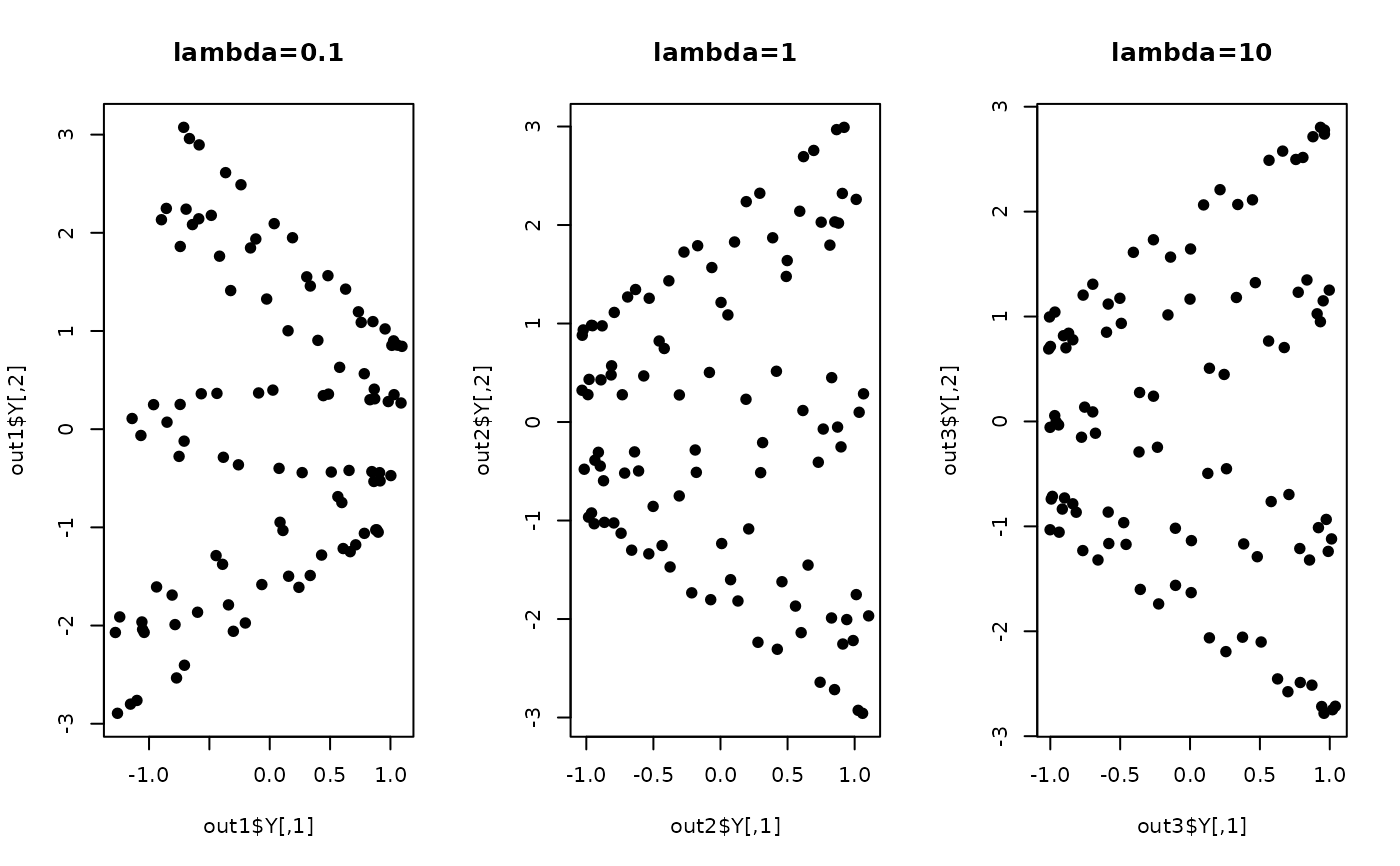
plot(out1$Y, pch=19, main="lambda=0.1")
plot(out2$Y, pch=19, main="lambda=1")
plot(out3$Y, pch=19, main="lambda=10")
 par(opar)
# }
par(opar)
# }